
Cresset Flare is a strong molecular modelling tool. It sounds complicated, but it’s fascinating. The software allows you to see the 3D structures of target proteins and possible ligands. Following that, you can examine their interactions and, finally, choose the most promising chemicals for future lab tests. I’m not sure whether anyone here is informed about this subject, but I thought I’d post it here in case anyone finds it beneficial.
Cresset Flare is available for free download via the link provided below. The archive includes an activation crack, but no key.
The author’s target audience consists mostly of computational and medicinal chemists working in pharmaceutical, biotech, and academic settings. They use Flare as a workflow: take a large number of candidate molecules, run them through various analysis tools, and end up with a small, curated set of the most promising variants, which they submit to the lab. The savings in time, resources, and reagents are enormous.
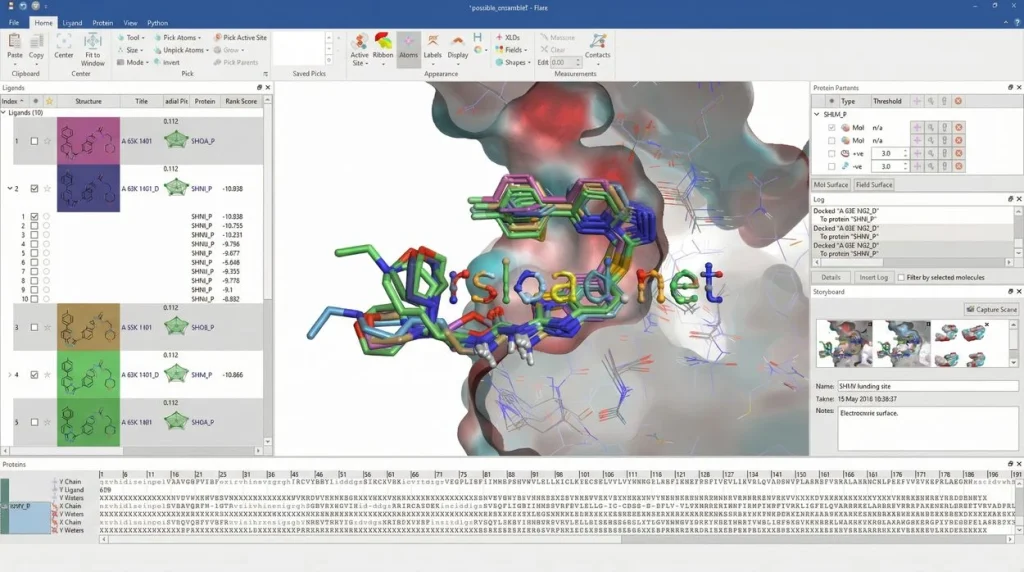
The program promotes two approaches to drug design:
- Ligand-based design involves comparing and prioritising molecules based on shape, electrostatics, and binding activity, and using QSAR models to predict the activity and ADMET properties of novel compounds.
- Docking and scoring, electrostatic complementarity, molecular dynamics, binding pocket analysis, and water analysis using GIST and 3D-RISM are all examples of structure-based design techniques.
Peculiarities:
- High-quality 3D visualisation of targets and ligands.
- Calculate perturbation-free energy to predict associated ligand activity accurately.
- Build predictive QSAR models for activity and ADME properties.
- Detailed investigation of structure-activity correlations
- Optimising lead connections using multiple factors.
- Integration with Torx to share data with colleagues at various phases of the DMTA cycle.
- binding pocket analysis and molecular dynamics


